Original Articles: 2025 Vol: 17 Issue: 1
In silico Drug Discovery of Natural Compounds Targeting LasI Signaling Protein in Quorum Sensing of Pseudomonas aeruginosa through Molecular Docking and Binding Interaction Studies
Muhammad Mudassar*, Raafia Ahmed Mughal
Department of Biosciences, Comsats University, Islamabad, Pakistan
*Corresponding Author:
Received: 14-Oct-2024, Manuscript No. JOCPR-24-150073; Editor assigned: 17-Oct-2024, PreQC No. JOCPR-24-150073 (PQ); Reviewed: 31-Oct-2024, QC No. JOCPR-24-150073; Revised: 15-Jan-2025, Manuscript No. JOCPR-24-150073 (R); Published: 22-Jan-2025, DOI:10.37532/0975-7384.2025.17(1).241.
Citation: Mudassar M, et al. 2025. In silico Drug Discovery of Natural Compounds Targeting LasI Signaling Protein in Quorum Sensing of Pseudomonas aeruginosa through Molecular Docking and Binding Interaction Studies. J. Chem Pharm. Res., 17:241.
Abstract
Pseudomonas aeruginosa is a type of bacteria that causes various infections in the human body, affecting organs like lungs and blood etc. Bacterial infections are highly prevalent and are usually treated with antibiotics. However, a significant concern with antibiotics is that they not only kill harmful bacteria but can also kill the beneficial ones. Pseudomonas aeruginosa uses a process called quorum sensing to communicate within their communities. The LasI protein is essential for producing a signaling molecule called N-(3-oxododecanoyl)-L-homoserine lactone (3O-C12- HSL) to carried out this communication. This study aims to use natural compounds to inhibit the LasI protein to stop the production of signaling molecules to disrupt quorum sensing. In this study molecular docking and structural validation of protein ligand complex were carried out to rank the 4000 natural compounds library from zinc database. Based on binding affinities ten potential compounds were selected. Their structural validation revealed mainly hydrophobic interactions with Ser, Val and Phe and hydrogen bonding. It also revealed that 50% of ligand covers the active site of target protein. Then top ten natural compounds were compared with already approved FDA drugs against target protein. Future more, binding affinity comparison showed that natural compounds have high binding affinities than already existing FDA drugs. In conclusion, natural compounds can act as a potential inhibitor against LasI protein to treat infections caused by pseudomonas aeruginosa.
Keywords
Quorum sensing; Pseudomonas aeruginosa; Targeting LasI signaling protein; Molecular docking; Binding interaction studies; Protein and ligand
Introduction
Quorum sensing serves as the fundamental communication process among bacteria within the same community. It involves the transmission of signals, known as auto inducers, facilitating cell-to-cell communication [1]. The production, detection, and response to these signaling molecules activate neighboring bacterial communities.
This process leads to the synthesis of biofilms, modulation of gene expression, and the production or release of specific proteins and toxins [2]. Pathogenic bacteria predominantly utilize quorum sensing, emphasizing their pivotal role in coordinating collective behaviors, such as biofilm formation and virulence factor expression [3]. In essence, quorum sensing enables bacteria to synchronize responses, contributing to their adaptability and virulence [4].
Pseudomonas aeruginosa is a versatile Gram-negative bacterium, recognized for its adaptability across various environments [5]. This opportunistic pathogen tends to cause infections in individuals with weakened immune systems or underlying health issues, often being associated with infections acquired in hospital settings. Quorum sensing is a communication mechanism employed by mostly every bacterium [6]. In its communication arsenal, Pseudomonas aeruginosa relies on three main quorum sensing systems, namely the las system, rhl system, and pqs system. Among these systems, the Las system takes center stage as the primary quorum sensing mechanism in Pseudomonas aeruginosa. The steps of quorum sensing in Pseudomonas aeruginosa include. P. aeruginosa produces small signaling molecules called autoinducers [7]. In the Las system, the primary autoinducer is N-(3- oxododecanoyl)-homoserine lactone (3-oxo-C12-HSL). The synthesis of autoinducers is typically catalyzed by specific enzymes, such as LasI in the case of the Las system. Once synthesized, the autoinducers are released into the surrounding environment [8]. As the P. aeruginosa population grows, the concentration of autoinducers in the environment increases proportionally. P. aeruginosa cells possess receptors, such as LasR, which specifically bind to the autoinducers. When an autoinducer binds to its corresponding receptor, it forms a complex that activates the LasR protein [9]. The activated LasR protein acts as a transcriptional regulator, influencing the expression of specific genes. In the case of P. aeruginosa, the Las system controls the expression of numerous virulence factors, including those involved in biofilm formation, secretion systems, and the production of toxins.
Materials and Methods
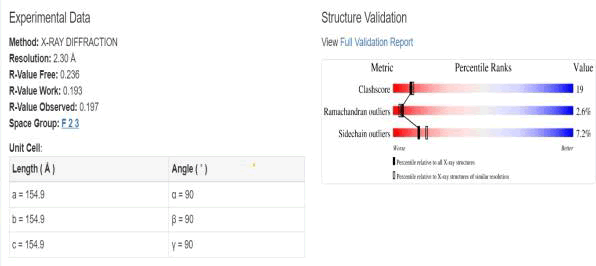
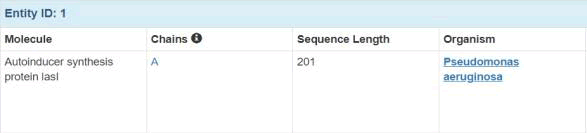
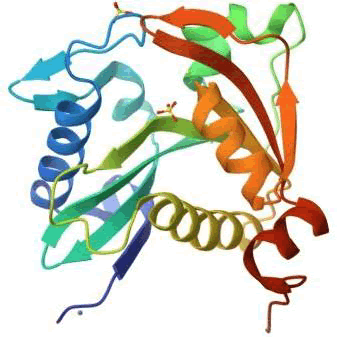
LasI is an essential signaling protein in Quorum sensing. It produces auto inducers which are necessary for Quorum sensing. The study uses 3D crystal structure of LasI protein obtained from protein data bank with ID number RO5. The protein's overall structure consists of an alternate (αβα) sandwich containing eight α-helices surrounded by nine β-strands. The active site is V-shaped, formed by a six-stranded beta sheet platform buttressed by three alpha helices, creating a substrate-binding cleft that leads to a tunnel accommodating the acyl-chain of acyl-ACP. This information is based on the crystal structure of the protein, as reported in various scientific articles and the Protein Data Bank. The structure was solved by X-Ray diffraction and structure was solved at 2.30 Å resolution. 1 chain(s) is present (1 unique chain) (Protein Data Bank) (Figure 1).
Figure 1: Experimental data.
LasI protein contain chain A which the presence of 201 amino acids. Taken from pseudomonas aeruginosa and having an expressing system of Escherichia coli. There is no mutation in system (Protein Data Bank) (Figures 2 and 3).
Figure 2: Sequence data.
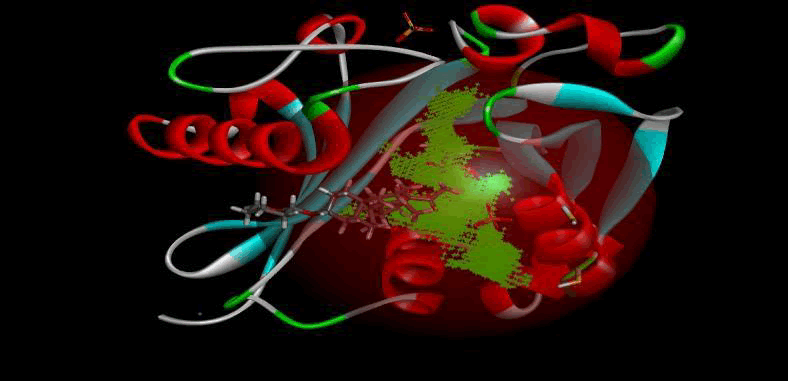
Figure 3: LasI protein crystal structure.
Ligand data contains 4000 natural compounds from different sources. 3120 natural compounds are taken from AnalytiCon discovery natural products (discovery). AnalytiCon company has excellence in natural product research and development for novel pharmaceutical, cosmetic and nutritional products. While the rest of the 880 natural compounds are taken from specs natural products (specs). Specs is the world's leading provider of compound management services and supplier of research (Zinc database) (Table 1).
| Name | Link to dataset |
| AnalytiCon natural products | https://zinc12.docking.org/catalogs/acdiscnp |
| Specs natural products | https://zinc12.docking.org/catalogs/specsnp |
Table 1: Products links to dataset.
Methods
The method includes the validation of protein structure and after that the molecular docking is used to check the binding affinity of ligand protein complex to check their stability.
Protein structure validation
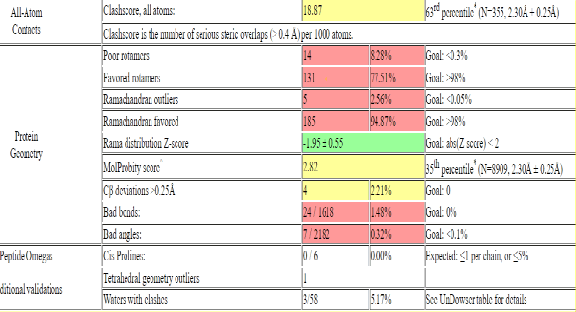
Protein structure validation, For the structure validation of LasI protein the Molprobity online tool is used. The structure is upload and the run is made to recheck the structure accuracy. The software identifies structure as LasI. Protein mainchain and sidechains are present. No explicit hydrogen atoms are included. 63 hetero group(s) are present. No explicit hydrogen atoms are included. 63 hetero group(s) is/are present. Refinement was carried out in REFMAC 5.1.24 i.e., =0.193; Rfree=0.236. For more validation click on continue to add hydrogen to protein structure. The result shows the molprobity score of 2.82 which is 35th percentile, Showing favorable score. Clash score is also favorable i.e., 18.87. The score suggested that the model has been refined to minimize steric clashes while Z score of 0.55 suggests that the distribution of torsional angles in protein structure is within an acceptable range. So this structure can be used for future study of protein ligand interaction (Figure 4).
Figure 4. Molprobity score.
Molprobity also shows the amino acids distribution in Ramachandran plot. Showing most of the amino acids in favorable area but close to warning area making protein a favorable structure to be used for future analysis (Figure 5).
Figure 5. Ramachandran plot.
Protein preparation, when preparing a protein structure for molecular docking using AutoDock Vina, a series of steps are undertaken to enhance the accuracy of the docking simulations. Initially, the three-dimensional structure of the target protein is loaded into the AutoDock Vina interface. To streamline the system and concentrate on relevant interactions, water molecules are removed from the structure, as they are often not directly involved in the binding events explored during docking. Subsequently, polar hydrogen atoms are added to the protein, ensuring a precise representation of hydrogen bonding interactions critical for molecular recognition. To facilitate the calculation of electrostatic interactions during docking simulations, Kollman charges are assigned to each atom in the protein structure. These partial charges are derived from a force field parameterization. Finally, the modified protein structure, now including atom charges and types, is saved in PDBQT format. This later enabling effective exploration of potential binding modes and interactions during molecular docking simulations (Figure 6).
Figure 6: Prepared protein structure for molecular docking.
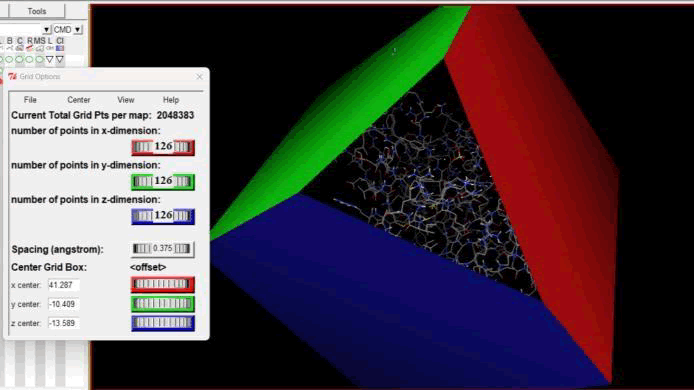
Following the preparation of the protein, the next step involves generating a grid box for the purpose of blind docking. This is achieved through the grid box section in Auto Dock Vina. Since the binding sites of the protein are unknown in this case, blind docking is employed, targeting the entire protein for potential ligand binding. The gird box is made with a spacing of 0.375 angstrom. The points in X dimension are 126 same as in y and z dimensions. The center grid box has values of X center =41.287, Y_center=-10.409 and Z_center=-13.589. Once the grid box is established, the parameters used for docking are saved, and the resulting information is stored in a text file. This file, containing the docking parameters, is renamed as "conf_vs txt" for clarity. The "conf_vs" designation suggests that these are configuration parameters related to blind docking against the entire protein structure (Figure 7).
Figure 7: Protein docking protocol.
Ligand preparation, to efficiently handle many ligands for molecular docking, the preparation process is streamlined using open babel software. This involves uploading the ligand dataset files individually in Structure Data File (SDF) format through the open babel interface. Once the datasets are loaded, the software automatically processes each ligand, converting them into separate files in PDBQT format. This automated conversion is a time-saving measure compared to manual preparation and ensures that each ligand is appropriately formatted for docking simulations. The output pathway is specified to organize the resulting PDBQT files, providing a systematic and structured approach to managing the prepared ligands for subsequent molecular docking studies.
Molecular docking
In molecular docking, the central step utilizes Perl software to dock 4000 ligands against a single protein, aiming to identify the ligand with the optimal binding affinity. As Autodock Vina's docking process for this large dataset is time-consuming, a Perl command is executed via the command line to expedite the procedure. To facilitate this, ligands, obtained through open babel, are consolidated in a folder along with the protein structure, Autodock Vina files, two critical text files (conf_vs. containing receptor information and ligand_names listing ligand names), and an accompanying Perl script. The Perl script, inherently part of the AutoDock Vina software, is integral to this process. This approach streamlines ligand evaluation, accelerates the identification of optimal binding candidates.
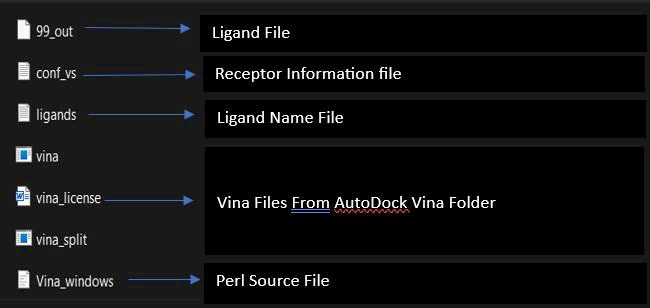
Following this, open the command line and enter the command
"dir/B>ligand.txt.", This command captures the names of all ligands and stores them in the ligand text file. It ensures that only ligands whose names are listed in this file will be subjected to the subsequent docking process. Next, execute the Perl script command:
"Perl Vina_windows.pl." This step initiates the docking process for all ligands against a single protein. The Perl script automatically generates log files and output pdbqt structures, providing detailed information about the docking interactions and results. After conducting molecular docking through Perl, ligands with higher binding affinities were prioritized, typically ranging near -6 and -5 kcal/mol. The top ten molecules with the lowest but higher affinities, ranging from -9 to -9.7 kcal/mol, were selected for further analysis. Among them, Zinc1995890 (Cedrelone) exhibited the highest binding affinity at -9.7 kcal/mol. Notable ligands in the top ten include ZINC01531236 (-9.5 kcal/mol), ZINC08382418 (3,21-Dihydroxypregnan-20-one, -9.4 kcal/mol), ZINC72320532 (-9.2 kcal/mol), ZINC04104769 (- 9.2 kcal/mol), ZINC04104770 (-9.2 kcal/mol), ZINC08382455 (Narcicrinine, -9 kcal/mol), ZINC04104771 (-9 kcal/mol), ZINC04104672 (-9 kcal/mol), and ZINC13298393 (-9 kcal/mol). These findings highlight promising candidates for further investigation based on their strong negative binding affinities in kcal/mol units (Table 2).
| Ligand no | Zinc ID | Binding affinity (kcal/mol) | Structure |
| 1 | Zinc19795 890 | -9.7 kcal/mol | 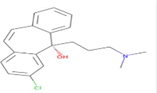 |
| 2 | Zinc19795 890 | -9.5 kcal/mol | 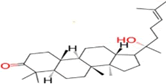 |
| 3 | Zinc19795 890 | -9.4 kcal/mol |  |
| 4 | Zinc19795 890 | -9.2 kcal/mol | 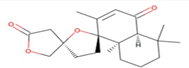 |
| 5 | Zinc19795 890 | -9.2 kcal/mol | 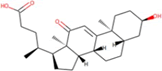 |
| 6 | Zinc19795 890 | -9.2 kcal/mol | 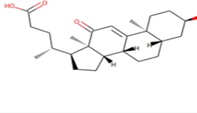 |
| 7 | Zinc19795 890 | -9 kcal/mol | 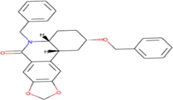 |
| 8 | Zinc19795 890 | -9 kcal/mol | 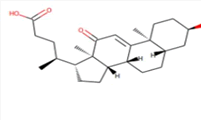 |
| 9 | Zinc19795 890 | -9 kcal/mol | 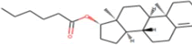 |
| 10 | Zinc19795 890 | -9 kcal/mol | 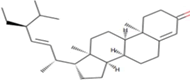 |
Table 2: Top molecule analysis.
After the selection of lowest binding affinity molecules are selected. After that the best poses are nominated which bind at lowest binding score with the active site of Protein. For this pymol used. The ligands are open in pymol then out of 9 poses select the pose with bind perfectly to the active site of protein (Table 3).
| Ligand number | Pose | RMSD value |
| Zinc1995890 | 1/9 | 0 |
| Zinc1531236 | 1/9 | 0 |
| Zinc0838418 | 1/9 | 0 |
| Zinc35455811 | 1/9 | 0 |
| Zinc04104769 | 1/9 | 0 |
Table 3: Values of the ligand number.
Binding sites validations
The study of binding site of protein plays a very important role in drug discovery. If the ligand in validation do not bind to the target binding, the desired pharmacological effect may not be achieved. This can lead to reduced drug efficacy. In the context of antimicrobial medicine, failure to bind at the target site can contribute to the development of drug resistance. For binding site validation Discovery Studio and Protein plus is used.
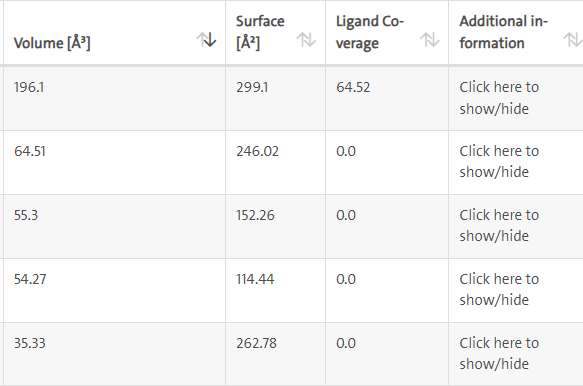
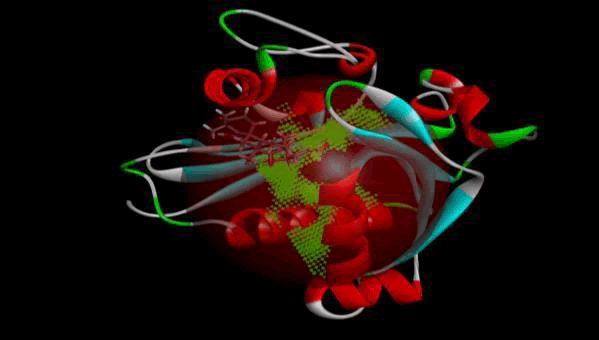
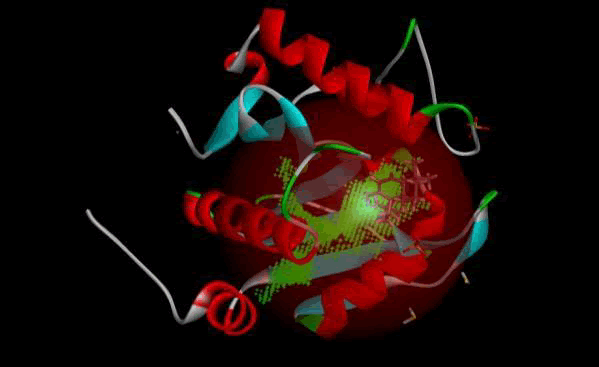
LasI Protein complex ligand Zinc1995890: LasI protein has five binding sites. The ligand binds at 1 binding sites with coverage of approximate 64.52%. The active site has the volume of 196.1 and surface of 299.1 (Figures 8 and 9).
Figure 8: Binding site score.
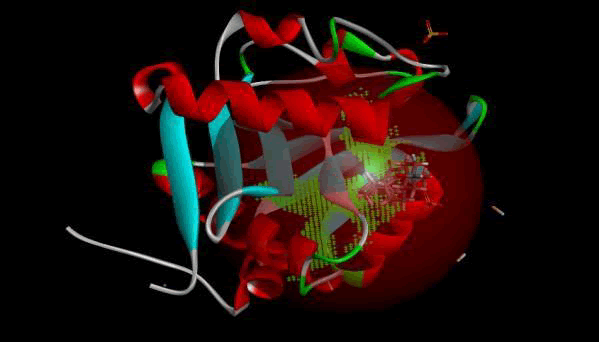
Figure 9: 3D view of active side.
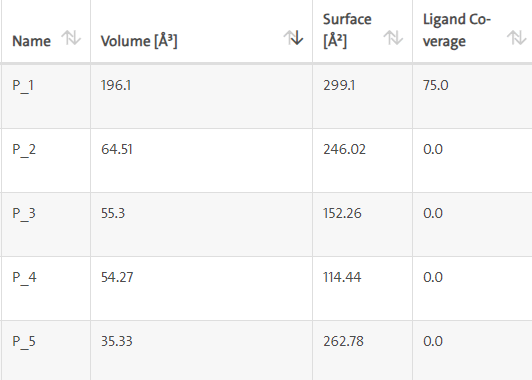
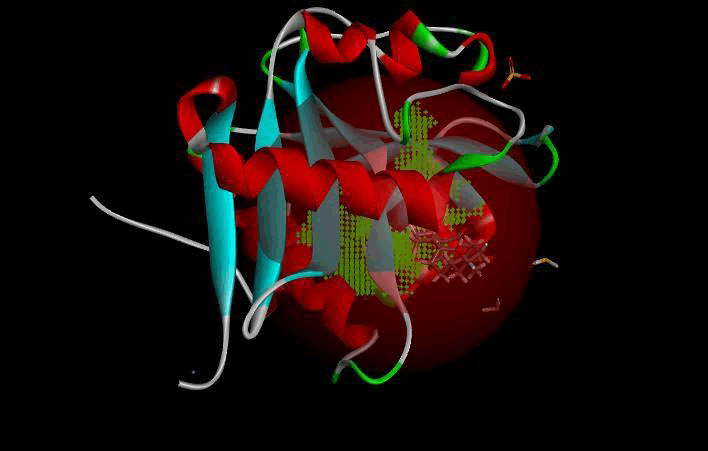
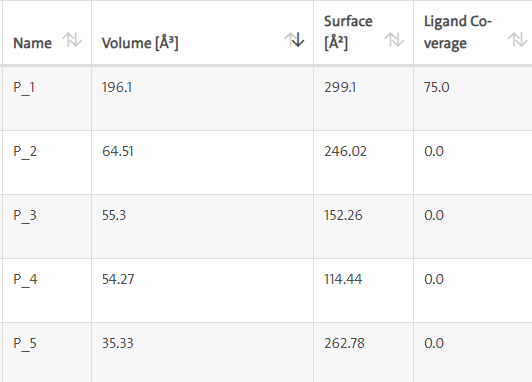
LasI protein complex with Zinc01531236: LasI protein has five binding sites. The ligand binds at 1 binding sites with coverage of approximate 75%. The active site has the volume of 196.1 and surface of 299.1 (Table 4 and Figures 10, 11).
| Zinc04104770 | 1/9 | 0 |
| Zinc72320532 | 1/9 | 0 |
| Zinc08382455 | 1/9 | 0 |
| Zinc04104672 | 1/9 | 0 |
| Zinc04104771 | 1/9 | 0 |
Table 4: Values of the ligand number.
Figure 10. Biding site score.
Figure 11: 3D view of active side.
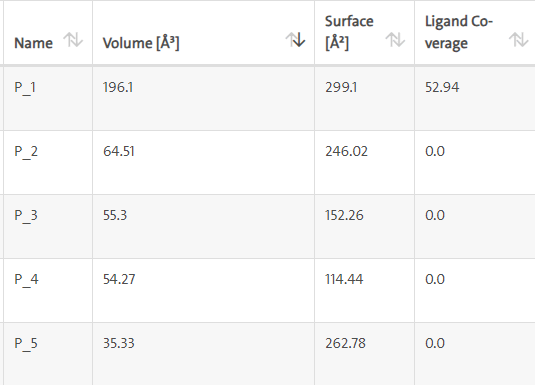
LasI protein complex with Zinc08382418: LasI protein has five binding sites. The ligand binds at 1st binding sites with coverage of approximate 52.94%. The active site has the volume of 196.1 and surface of 299.1 (Figures 12 and 13).
Figure 12: Binding site score.
Figure 13: 3D view of active side.
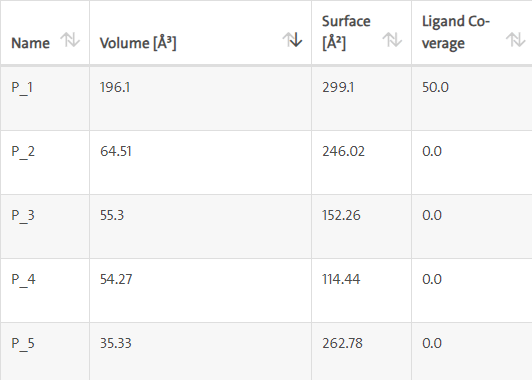
LasI protein complex with Zinc35455811: LasI protein have five binding sites. The ligand binds at 1st binding sites with coverage of approximate 50%. The active site has the volume of 196.1 and surface of 299.1 (Figures 14 and 15).
Figure 14: Binding site score.
Figure 15: 3D view of active side.
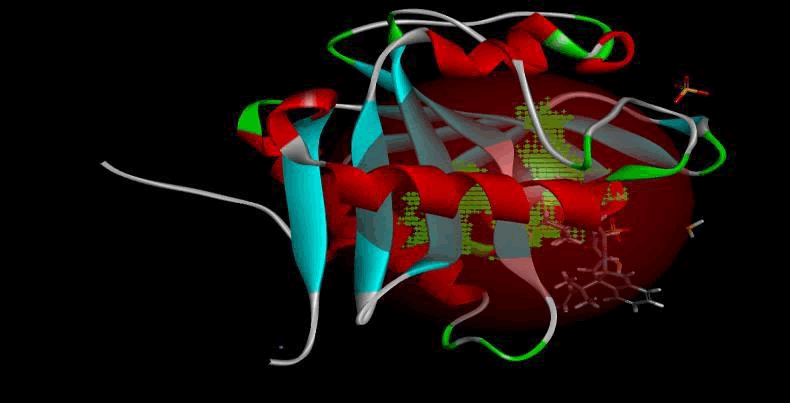
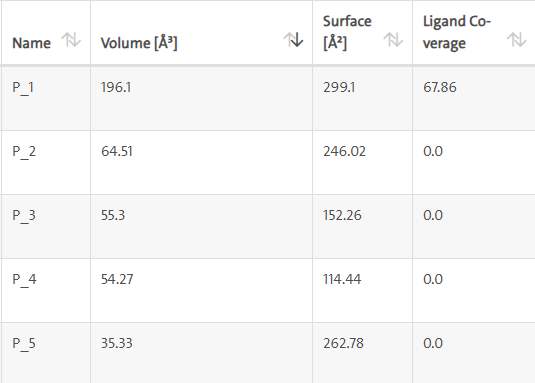
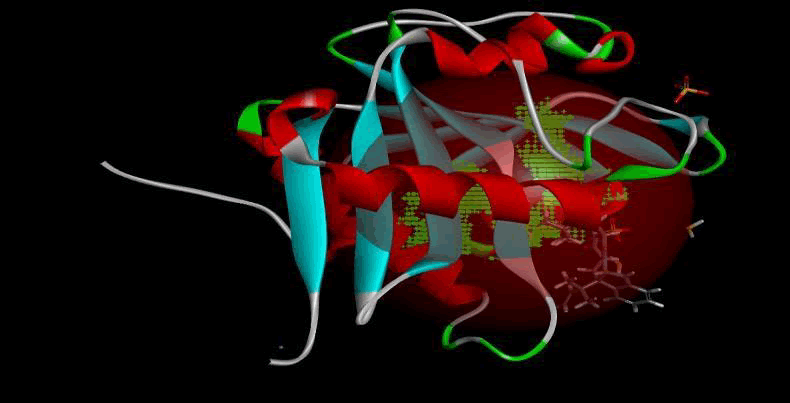
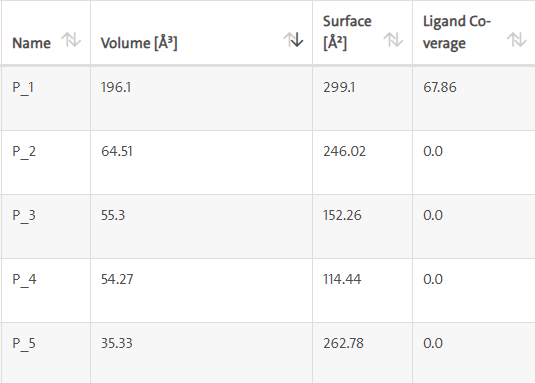
LasI protein and Zinc04104769: LasI protein has five binding sites. The ligand binds at 1st binding sites with coverage of approximate 67.86%. The active site has the volume of 196.1 and surface of 299.1 (Figures 16 and 17).
Figure 16: Binding site score.
Figure 17: 3D view of active side.
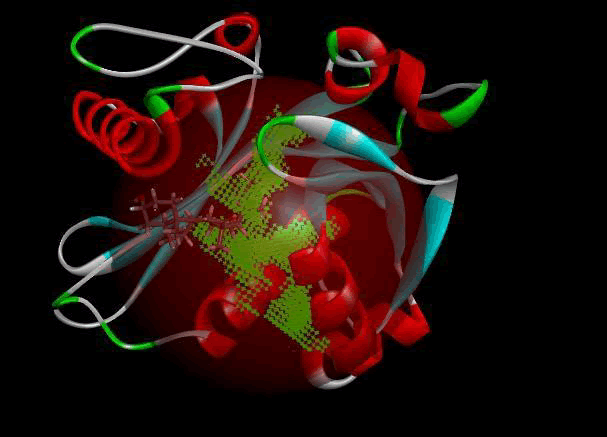
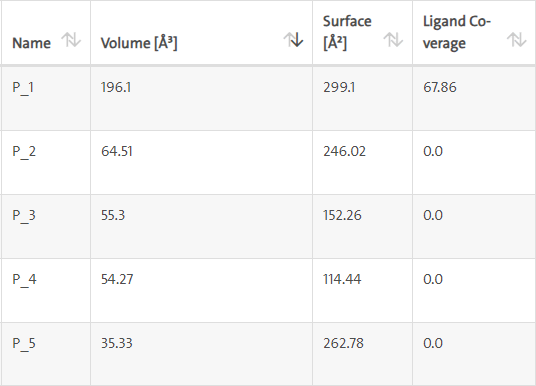
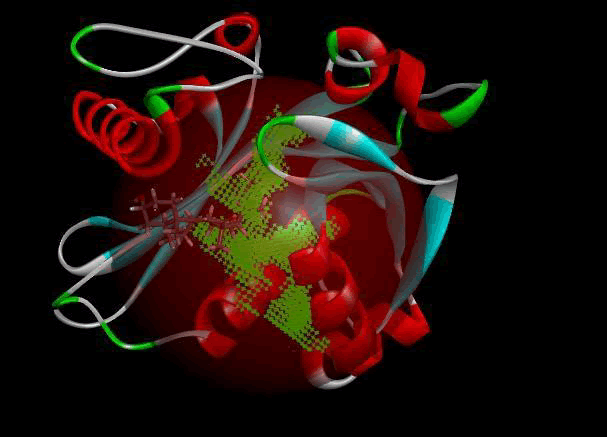
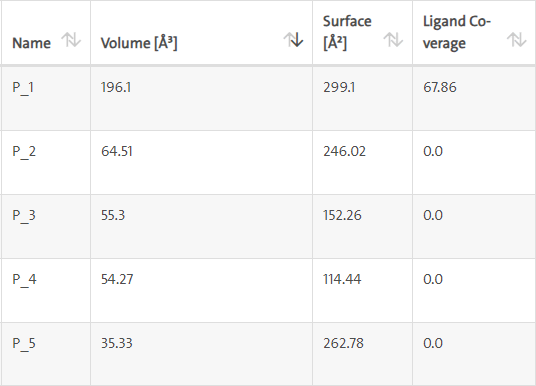
LasI protein complex with Zinc04104770: The protein has five binding sites. The ligand binds at 1st binding sites with coverage of approximate 67.86%. The active site has a volume of 196.1 and surface of 299.1 (Figures 18 and 19).
Figure 18: Binding site score.
Figure 19: 3D view of active side.
LasI protein and Zinc04104769: LasI protein has five binding sites. The ligand binds at 1st binding sites with coverage of approximate 67.86%. The active site has the volume of 196.1 and surface of 299.1 (Figures 20 and 21).
Figure 20: Binding site score.
Figure 21: 3D view of active side.
LasI protein complex with Zinc04104770: The protein has five binding sites. The ligand binds at 1st binding sites with coverage of approximate 67.86%. The active site has a volume of 196.1 and surface of 299.1 (Figures 22 and 23).
Figure 22: Binding site score.
Figure 23: 3D view of active side.
LasI protein complex with Zinc72320532: LasI protein have five binding sites. The ligand binds at 1st binding sites with coverage of approximate 75%. The active site has the volume of 196.1 and surface of 299.1 (Figures 24 and 25).
Figure 24: Binding site score.
Figure 25: 3D view of active side.
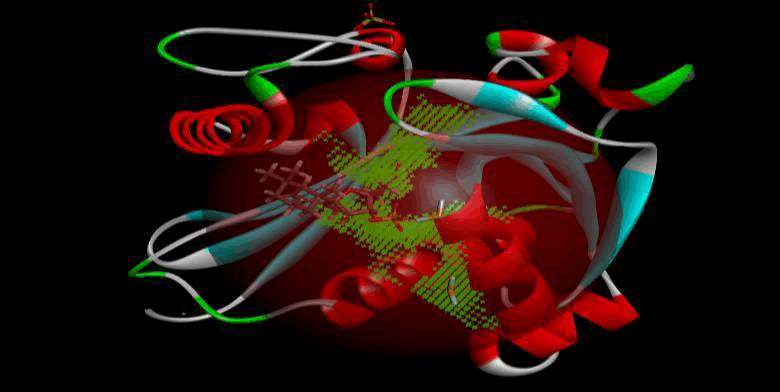
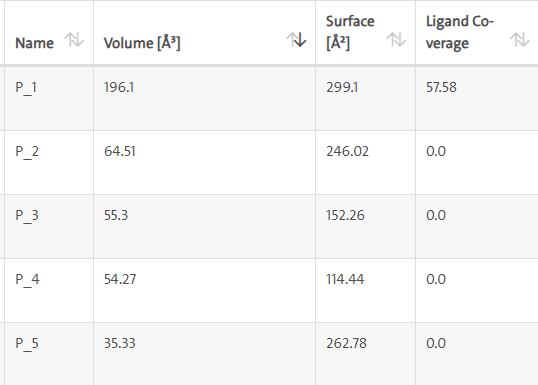
LasI protein Complex with Zinc08382455: LasI protein has five binding sites. The ligand binds at 1st binding sites with coverage of approximate 57.58%. The active site has the volume of 196.1 and surface of 299.1 (Figures 26 and 27).
Figure 26: Binding site score.
Figure 27: 3D view of active side.
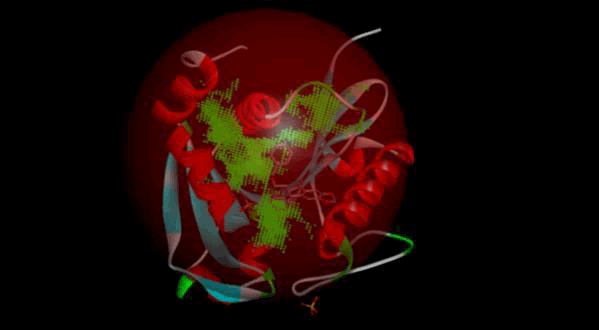
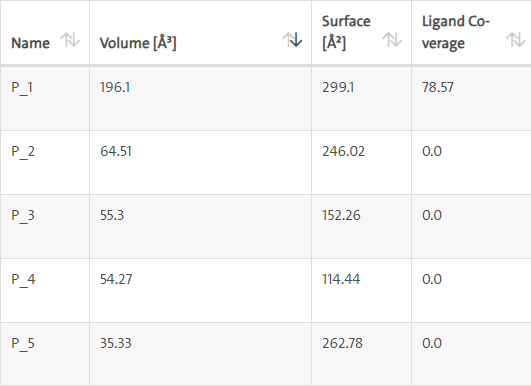
LasI protein complex with Zinc04104672: The protein has five binding sites. The ligand binds at 1st binding sites with coverage of approximate 78.57%. The active site has the volume of 196.1 and surface of 299.1 (Figures 28 and 29).
Figure 28: Binding site score.
Figure 29: 3D view of active side.
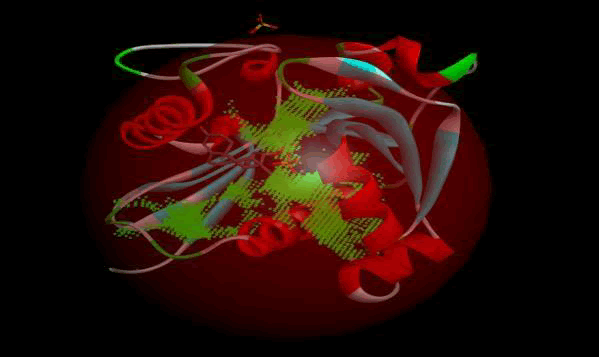
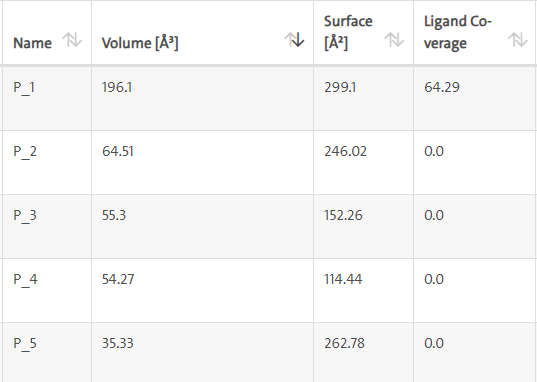
LasI protein complex with Zinc72320532: The protein has five binding sites. The ligand binds at 1st binding sites with coverage of approximate 64.29%. The active site has the volume of 196.1 and surface of 299.1 (Figures 30 and 31).
Figure 30: Binding site score.
Figure 31: 3D view of active side.
Validation of hydrogen bonds and pi-pi interaction
Validating hydrogen bonds and pi-pi interaction in a protein-ligand complex is crucial for decoding molecular recognition intricacies. These interactions govern binding specificity and strength, profoundly impacting complex stability and functionality. This validation process is pivotal for advancing our comprehension of molecular mechanisms, specifically in drug repurposing. By unraveling these intricacies, we gain unique insights to explore therapeutic avenues, optimize repurposing strategies, and drive innovative advancements in medical research.
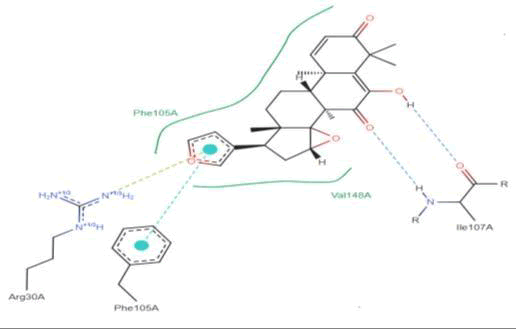
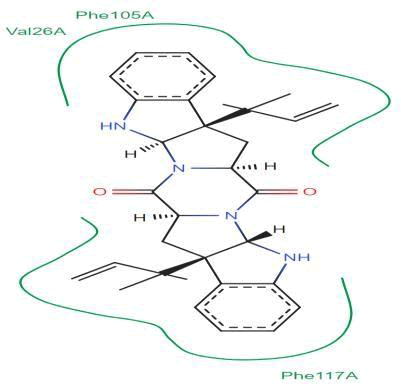
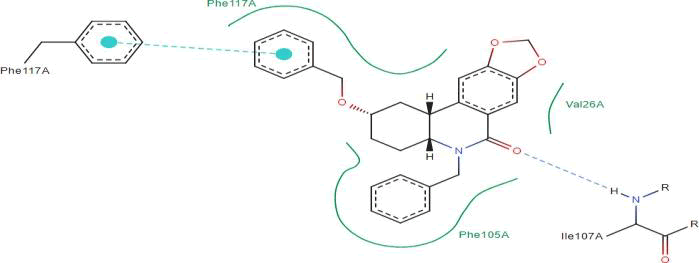
LasI Protein complex ligand Zinc1995890: In LasI Protein complex ligand Zinc1995890, two hydrogen bonds are present with one pi-pi interaction. The 2D diagram shows blue line as hydrogen bond, red as pi-pi interaction and green has hydrophobicity. The hydrogen bonds are between ligand and ILE107A were ligand act as acceptor and ILE act as donor. The pi- pi interactions are between ligand and PHE105A (Figure 32).
Figure 32: 2D diagram of interactions.
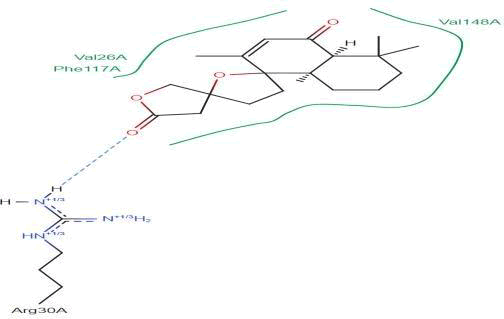
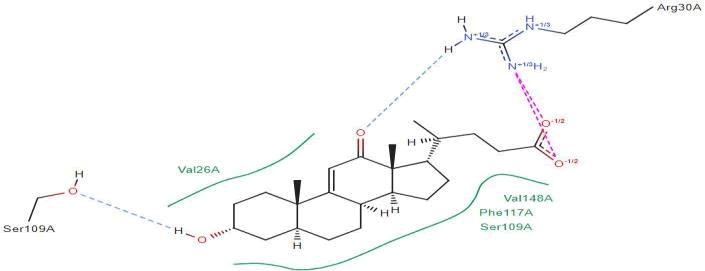
LasI Protein complex ligand Zinc01531236: In LasI Protein complex ligand Zinc01531236, one hydrogen bond is present with no pi-pi interaction. The 2D diagram shows the blue line as hydrogen bond. The hydrogen bond is between ligand and ARG30A, where ligand acts as acceptor and Arg30A acts as donor (Figure 33).
Figure 33: 2D diagram of interactions.
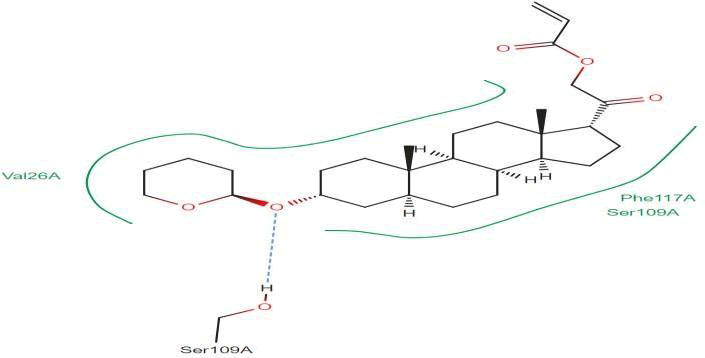
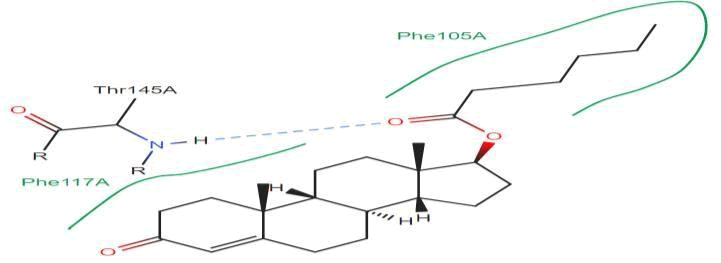
LasI Protein complex ligand Zinc08382418: In LasI Protein complex ligand Zinc08382418, one hydrogen bond is present with no pi-pi interaction. The 2D diagram shows the blue line as hydrogen bond. The hydrogen bond is between ligand and Ser109A, where ligand acts as acceptor and Ser109A acts as donor (Figure 34).
Figure 34: 2D diagram of interactions.
LasI Protein complex ligand Zinc35455811: In LasI Protein complex ligand Zinc35455811, there is no hydrogen bond and no pi-pi interaction (Figure 35).
Figure 35: 2D diagram of interactions.
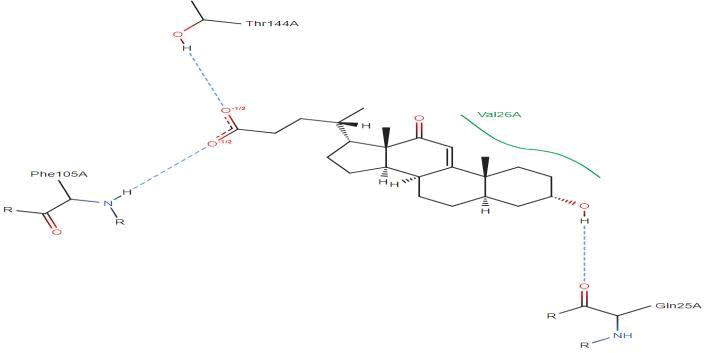
LasI Protein complex ligand Zinc04104769: In LasI Protein complex ligand Zinc04104769, three hydrogen bonds are present with no pi-pi interaction. The one hydrogen bond is between ligand and Phe105A. Second with Thr144A and last with Gln25A (Figure 36).
Figure 36: 2D diagram of interactions.
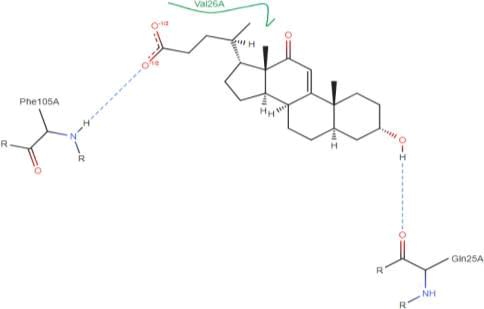
LasI Protein complex ligand Zinc04104770: In LasI Protein complex ligand Zinc04104770, two hydrogen bonds are present with no pi-pi interaction. The one hydrogen bond is between ligand and Phe105A and Second with Gln25A (Figure 37).
Figure 37: 2D diagram of interactions.
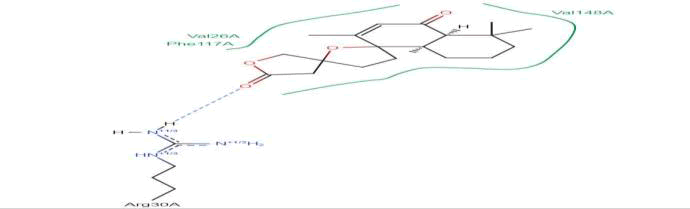
LasI Protein complex ligand Zinc72320532: In LasI Protein complex ligand Zinc72320532, one hydrogen bond is present with no pi-pi interaction. The hydrogen bond is between ligand and Arg30A, where ligand acts as acceptor and Arg30A acts as donor (Figure 38).
Figure 38: 2D diagram of interactions.
LasI Protein complex ligand Zinc08382455: In LasI Protein complex ligand Zinc08382455, one hydrogen bond is present with one pi-pi interaction. The hydrogen bond is between ligand and Ile107A, where ligand acts as acceptor and Ile107A act as donor. Pi-pi interaction is between ligand and phe 117A (Figure 39).
Figure 39: 2D diagram of interactions.
LasI Protein complex ligand Zinc04104672: In LasI Protein complex ligand Zinc04104672, one hydrogen bond present with no pi-pi interaction. The one hydrogen bond is between ligand and Phe117A, where ligand acts as acceptor and phe117A act as donor (Figure 40).
Figure 40: 2D diagram of interactions.
LasI Protein complex ligand Zinc04104771: In LasI Protein complex ligand 04104771, two hydrogen bonds are present with no pi-pi interaction. The one hydrogen bond is between ligand and Ser109A, where ligand acts as acceptor and Ser109A act as donor. Second with Arg30A where ligand act as acceptor and Arg30A as donor (Figure 41).
Figure 41: 2D diagram of interactions.
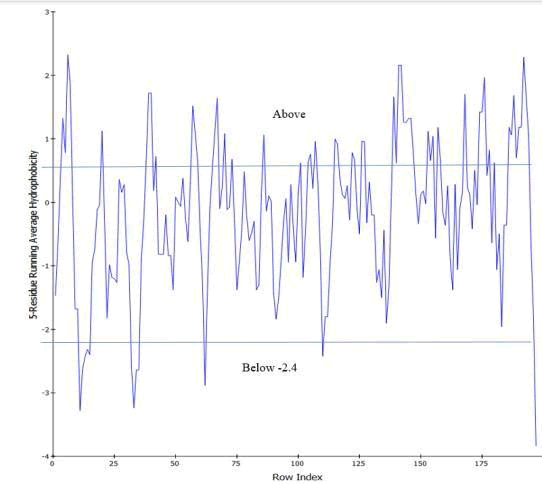
Protein hydrophobicity is viewed using discovery studio. In discovery studio hydrophobicity chart is used to view the overall nature of protein. The hydrophobicity scale is taken from Kyte and Doolittle. Residues with a hydrophobicity of 0.7 or more are hydrophobic and those under -2.4 are hydrophilic. Keep that in view and the chart appearance shows that mostly residues are in between 0.7 and -2.4 which is neutral. After that mostly are hydrophobic (Figure 42).
Figure 42: Hydropathy plot showing 5-residue running average of hydrophobicity.
Among the selected ten ligands for further study, hydrophobic interactions vary based on distinct amino acid residues. For instance, in Zinc1995890, hydrophobic interactions are instigated by Phe105 and Val148. In Zinc01531236, Val26A, Val148, and Phe117 contribute to hydrophobic interactions. Similarly, Zinc08382418 features hydrophobic interactions driven by Val26A, Ser109A, and Phe117A. In Zinc35455811, Val26A, Phe105A, and Phe117 are involved in hydrophobic interactions. Val26 is the key contributor in Zinc04104769 and Zinc04104770. Hydrophobic interactions in Zinc72320532 are influenced by Val26, Val148, and Phe117. In Zinc08382455, Val26, Phe105, and Phe117 contribute to hydrophobic interactions. Zinc04104672 showcases hydrophobic interactions mediated by Phe105 and Phe117. Lastly, Zinc04104771, Val26A, Val148, Phe117, and Ser109A are responsible for hydrophobic interactions (Table 5).
|
Ligands name |
Amino acid |
Residues number |
|
Zinc1995890 |
Phe |
105 |
|
In Zinc01531236 |
Val |
26 |
|
Zinc08382418 |
Val |
26 |
|
Zinc35455811 |
Val |
26 |
|
Zinc04104769 |
Val |
26 |
|
Zinc04104770 |
Val |
26 |
|
Zinc72320532 |
Val |
26 |
|
Zinc08382455 |
Val |
26 |
|
Zinc04104672 |
Phe |
105 |
|
Zinc04104771 |
Val |
Val148 |
Table 5: Interaction of ligands with key amino acid residues in the target protein.
Comparing with already FDA approved drugs
For these five already approved drugs(inhibitors) of LasI signaling protein are taken and their structure are retrieved from PubChem. Then the already prepared LasI protein for the docking is taken, by using pearl script these 5 FDA approved drugs are taken and are dock against LasI protein to check their binding affinity. The binding affinity for these % drugs are between 4.2 to 5.7 (Table 6).
| Ligand name | Lowest binding affinity | RMSD value |
| Carbapenems | -4.7 | 0 |
| Fosfomycin | -4.2 | 0 |
| Chalcone | -5.7 | 0 |
| Furanone C-30 | -5 | 0 |
| Colistina | -5.2 | 0 |
Table 6: Binding affinities and RMSD values of selected ligands.
Results
Traditional antibiotics are the major challenge of today, as they have many negative impacts on human health. Much research says that the Traditional antibiotics not only killed harmful bacteria but also the bacteria which are useful for the body. For that, this study aimed to explore the potential of natural compounds as inhibitors against the LasI signaling protein, a crucial component in Pseudomonas aeruginosa quorum sensing. The method of molecular docking and structural validations are applied to check the protein ligand complexes.
Interpretation of molecular docking results:
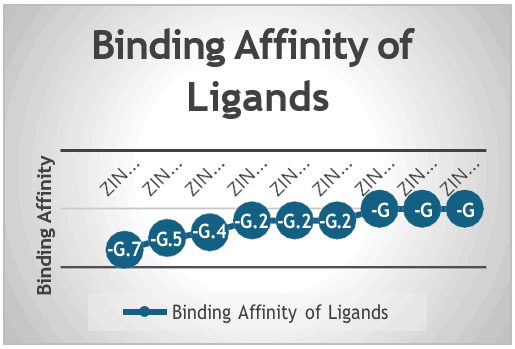
The molecular docking results provide valuable insights into the potential of natural compounds as inhibitors against the LasI signaling protein. 4000 natural products are dock against LasI protein, which is very important protein in quorum sensing. The docking results show the average docking score of 6 kcal/mol and 7 kcal/mol. But after the validation of all 4000 molecules, 10 molecules with the lowest binding affinity are selected. The result of these ten molecules shows that Zinc1995890 (Cedrelone) exhibited the highest binding affinity at -9.7 kcal/mol while ZINC01531236 has -9.5 kcal/mol, ZINC08382418 (3,21-Dihydroxypregnan-20-one) has -9.4 kcal/mol, ZINC72320532 has -9.2 kcal/mol, ZINC04104769 has -9.2 kcal/mol, ZINC04104770 has -9.2 kcal/mol, ZINC08382455 (Narcicrinine) has -9 kcal/mol, ZINC04104771 has -9 kcal/mol, ZINC04104672 has -9 kcal/mol, and ZINC13298393 has -9 kcal/mol. The high binding affinity of Cedrelone suggests a strong interaction with the protein, possibly disrupting its quorum sensing function. The consistent negative binding affinities across multiple ligands reinforce the hypothesis that natural products can effectively modulate LasI protein activity (Figure 43).
Figure 43: Graph depicting binding affinity of ligands from ZINC database.
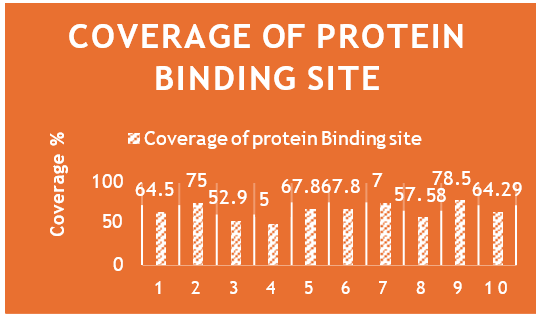
Structural validation and ligand binding sites: The validation of the LasI protein structure ensures the reliability of the docking results. The MolProbity analysis and Ramachandran plot confirm the structural integrity of the protein, supporting its suitability for ligand interaction studies. The Molprobity score are positive showing a good protein structure for future study. The identification of specific binding sites for each ligand provides a foundation for understanding the diverse interactions between natural compounds and the LasI protein. Each pose of ligand which is selected is view and check for binding score. The ligand coverage shows that all the ten molecules cover the protein binding sites more than 50% showing it a stable complex. The highest ligand coverage is done by Zinc04104672 of 78% while the second highest coverage is done by Zinc01531236 and Zinc72320532 (Figure 44).
Figure 44: Coverage percentage of protein binding sites across multiple ligands.
The analysis of hydrogen bonds, pi-pi interactions, and hydrophobic contacts sheds light on the molecular mechanisms governing the stability of ligand-protein complexes. Approximately, all molecule form hydrogen bond while there is only one ligand which do not form any hydrogen bond. Cedrelone which have highest binding affinity forms hydrogen bonds and pi-pi interactions, indicating specific amino acid residues crucial for binding. Hydrophobic contacts further contribute to the overall stability of the complexes, emphasizing the multifaceted nature of the interactions. The hydrophobicity chart shows that protein mostly residues are neutral, but some are hydrophobic. The hydrophobic interactions are mostly created by ILE, SER, VAL, and PHE. Phenylalanine, with its aromatic ring, tends to exhibit stronger hydrophobic interactions as compared to other but other also can involve in hydrophobic interactions.
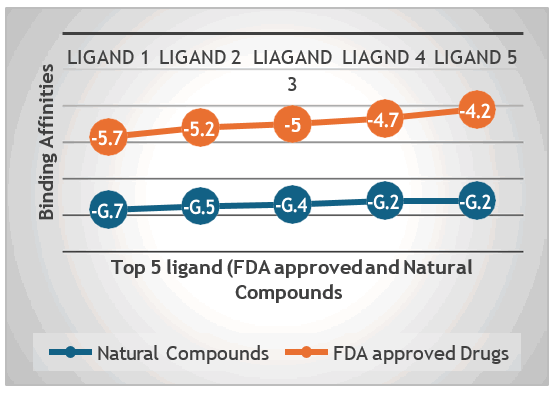
Comparison with FDA-approved drugs: A comparison with FDA-approved drugs known as LasI protein inhibitors revealed that the top ligands from the study demonstrated binding affinities comparable to these existing inhibitors. The binding affinity of FDA approved drugs falls between the range of -4 kcal/mol to 5.7 kcal/mol while comparing them with the selected top ten natural product those binding affinity range from 9 kcal/mol to 9.7 kcal/mol. This finding suggested that natural compounds could rival conventional drugs in their potential to inhibit LasI signaling protein (Figure 45).
Figure 45: Comparison of binding affinities between natural compounds and FDA-approved drugs.
Discussion
The findings of this study present compelling evidence supporting the potential of natural compounds as inhibitors against the LasI signaling protein, a critical component in Pseudomonas aeruginosa quorum sensing. The molecular docking results revealed that out of 4000 natural products, Cedrelone (Zinc1995890) exhibited the highest binding affinity at -9.7 kcal/mol. This suggests a strong interaction with the LasI protein, potentially disrupting its quorum sensing function. The consistent negative binding affinities across multiple ligands reinforce the hypothesis that natural products can effectively modulate LasI protein activity.
Structural validation using MolProbity analysis and Ramachandran plot confirmed the reliability of the LasI protein structure for ligand interaction studies. The identification of specific binding sites for each of the top ten ligands, along with ligand coverage exceeding 50%, indicates stable and reliable complexes. Cedrelone, with its high binding affinity, also demonstrated a considerable coverage of the protein binding site.
Analysis of hydrogen bonds, pi-pi interactions, and hydrophobic contacts provided insights into the molecular mechanisms governing the stability of ligand-protein complexes. Cedrelone, with its highest binding affinity, formed hydrogen bonds and pi-pi interactions, emphasizing specific amino acid residues crucial for binding. Hydrophobic interactions, predominantly involving ILE, SER, VAL, and PHE, contributed to the overall stability of the complexes.
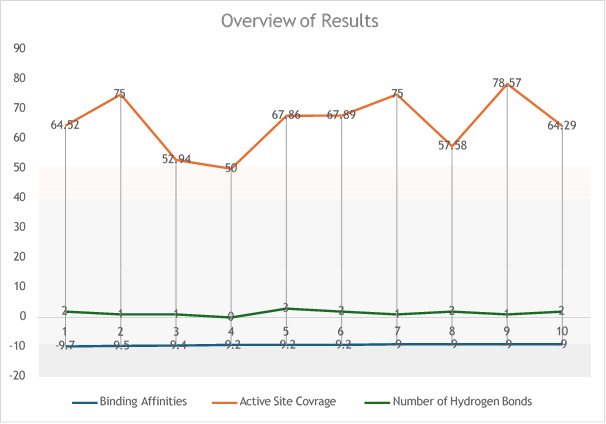
Comparison with FDA-approved drugs known as LasI protein inhibitors showed that the top ligands from the study exhibited binding affinities comparable to existing inhibitors. The natural compounds surpassed the binding affinity range of FDA-approved drugs, ranging from 9 kcal/mol to 9.7 kcal/mol, suggesting that these natural compounds could rival conventional drugs in inhibiting LasI signaling protein (Figure 46).
Figure 46: Comprehensive overview of ligand-protein interaction parameters.
Conclusion
The results aligned with the overarching hypothesis that natural compounds could serve as effective inhibitors against the LasI protein. Cedrelone emerged as a standout candidate with the highest binding affinity. Other factors include binding site coverage, formation of hydrogen bond, hydrophobic interactions also suggest it as a good inhibitor of LasI protein. The other natural products also show a good binding affinity against LasI protein as compared to conventional medicine. Their interactions, hydrogen bonding also show that they can be good inhibitors od the LasI protein. So, after the analysis of natural compounds using molecular docking d structure validation, we can say that natural products can be a promising new drug in the treatment of infections caused by the pseudomonas aeruginosa and against the signaling molecules produce against the Quorum sensing.
References
- Atanasov AG, et al. Nat Rev Drug Discov. 2021;20(3):200-216.
[Crossref] [Google Scholar] [PubMed]
- Pesci EC, et al. J Bacteriol. 1997;179(10):3127-32.
[Crossref] [Google Scholar] [PubMed]
- Guzzo F, et al. Molecules. 2020;25(21):5024.
[Crossref] [Google Scholar] [PubMed]
- Haoran Li, et al. Mar Drugs. 2022;20(3):192.
[Crossref] [Google Scholar] [PubMed]
- de Kievit TR. Environ Microbiol. 2009;11(2):279-288.
[Crossref] [Google Scholar] [PubMed]
- Zhong L, et al. Int J Mol Sci. 2020;21(6):2190.
[Crossref] [Google Scholar] [PubMed]
- O'Boyle NM, et al. J Cheminform. 2011;3:1-4.
[Crossref] [Google Scholar] [PubMed]
- Qin S, et al. Signal Transduct Target Ther. 2022;7(1):199.
[Crossref] [Google Scholar] [PubMed]
- Spagnolo AM, et al. Rev Res Med Microbiol. 2021;32(3):169-175.
- Rutherford ST, et al. Cold Spring Harb Perspect Med. 2012;2(11):a012427.
[Crossref] [Google Scholar] [PubMed]