Original Articles: 2025 Vol: 17 Issue: 1
Amide Linkages: Strategies to Combat Antibiotic Resistance
Vidya Sagar Jerra*
Department of Chemistry, School of Applied Sciences and Humanities, Vignan’s Foundation for Science, Technology and Research, Vadlamudi, India
*Corresponding Author:
- Vidya Sagar Jerra Department of Chemistry, School of Applied Sciences and Humanities, Vignan’s Foundation for Science, Technology and Research, Vadlamudi, India
Received: 03-Oct-2024, Manuscript No. JOCPR-24-149468; Editor assigned: 07-Oct-2024, PreQC No. JOCPR-24-149468 (PQ); Reviewed: 21-Oct-2024, QC No. JOCPR-24-149468; Revised: 15-Jan2025, Manuscript No. JOCPR-24-149468 (R); Published: 22-Jan-2025, DOI:10.37532/0975-7384.2025.17(1).240.
Citation: Jerra VS. 2025. Amide Linkages: Strategies to Combat Antibiotic Resistance. J. Chem Pharm. Res., 17:240.
Abstract
This review paper highlights the pivotal role of amide linkages in overcoming the limitations of traditional antibiotics and discusses recent progress in Antimicrobial Peptide (AMP) design. Amide linkages play a crucial role in enhancing the stability, bioavailability, and efficacy of AMPs, particularly in addressing the variable effectiveness against Gram-positive and Gram-negative bacteria. A thorough understanding of amide linkage dynamics is essential for developing next-generation AMP-based therapeutics, offering a viable solution to the escalating issue of antibiotic resistance. By optimizing amide bonds, along with incorporating non-natural amino acids and peptide cyclization strategies, researchers are making significant strides in overcoming challenges such as proteolytic degradation, toxicity, and bacterial resistance. These advancements, combined with innovations like peptide mimetics and PEGylation, are helping to unlock the full clinical potential of peptide antibiotics. Continued efforts in refining Structure-Activity Relationships (SAR) and monitoring bacterial resistance mechanisms will be key to translating AMP-based therapeutics into viable clinical treatments and strengthening the global fight against multidrug-resistant infections.
Keywords
Peptide antibiotics; Antimicrobial Peptides (AMPs); Amide linkages; Antibiotic resistance; Proteolytic stability; Bioavailability; Peptide cyclization; Non-natural amino acids; Peptide mimetics; PEGylation; Multidrug- resistant infections; Structure Activity Relationships (SAR); Peptide therapeutics
Introduction
The escalating crisis of antibiotic resistance poses a severe threat to global health, as resistant pathogens like Methicillin-Resistant Staphylococcus aureus (MRSA) and Carbapenem-Resistant Enterobacteriaceae (CRE) undermine the efficacy of traditional antibiotics [1]. According to recent reviews, the factors driving this crisis include the overuse and misuse of antibiotics, which accelerate resistance mechanisms such as enzyme-mediated degradation, altered target sites, and biofilm formation [2]. Current trends in antibacterial drug discovery are focusing on innovative approaches to combat these challenges [3]. Strategies under exploration include bacteriophage therapy, CRISPR-based antimicrobials, and novel small molecules targeting unexploited bacterial pathways [4]. Despite their potential, these methods often face hurdles related to scalability, delivery, and efficacy [5].
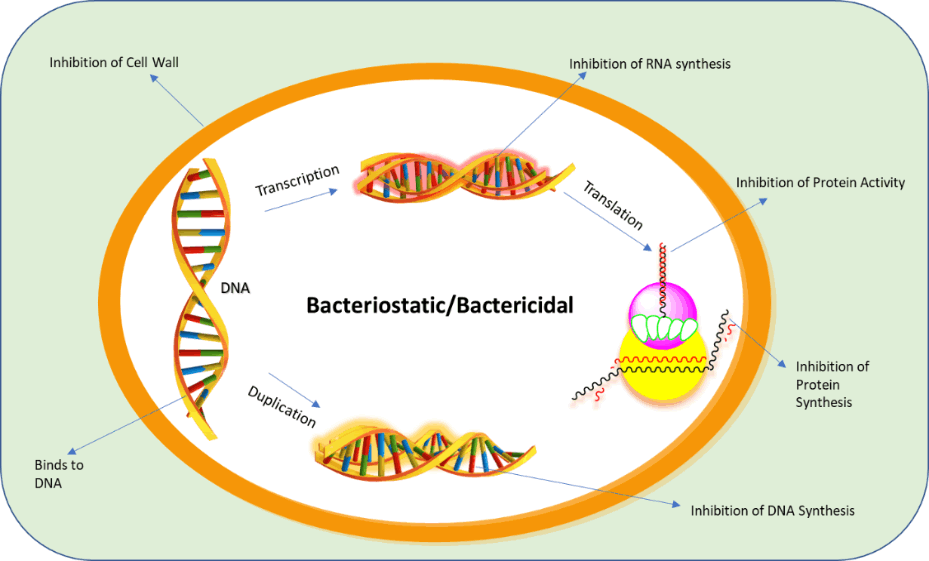
In the ongoing search for new antibacterial agents, peptide antibiotics are often lauded for their broad-spectrum activity against diverse pathogens [6]. However, achieving consistent potency across different bacterial species presents significant challenges [7]. For instance, while some peptides may exhibit strong activity against Gram-positive bacteria, their effectiveness can be limited against Gram-negative bacteria due to the latter’s protective outer membrane [8]. This variability underscores the need for targeted modifications to enhance peptide efficacy across a wider range of bacterial types (Figure 1) [9].
Figure 1: Targeted modifications to enhance peptide efficacy across a wider range.
Addressing these challenges-such as the complexity of synthesis, stability issues, toxicity, understanding Structure-Activity Relationships (SAR), resistance development, and achieving broad-spectrum activity-is crucial for fully realizing the potential of peptide-derived antibiotics [10]. Recent efforts have focused on several key areas to overcome these obstacles, particularly in relation to amide linkages and overall peptide stability [11]. Key issues include overcoming bacterial resistance, enhancing stability against proteolysis, improving bioavailability, minimizing host cell toxicity, and scaling up production for clinical applications [12].
Antimicrobial Peptides (AMPs) offer a compelling solution to the antibiotic resistance crisis [13]. AMPs are short peptides that exhibit broad-spectrum activity against bacteria, fungi, and viruses by disrupting microbial membranes and inhibiting intracellular targets [14]. They also possess immune-modulatory properties, enhancing the host's ability to fight infections [15]. Recent research underscores the importance of AMPs in addressing antibiotic resistance due to their unique mechanisms of action that minimize the likelihood of resistance development [16].
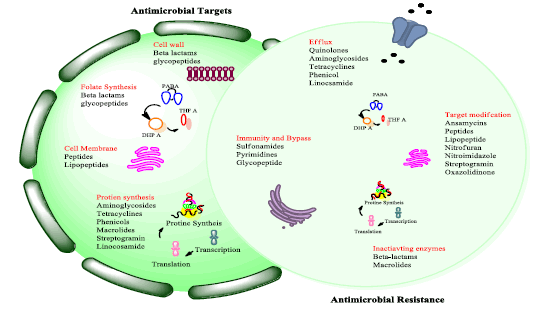
Recent advancements in AMP design, such as modifications to amide linkages, incorporation of non-natural amino acids, and cyclization techniques, have significantly enhanced their stability, selectivity, and therapeutic potential [17]. These innovations have led to improved resistance to proteolytic degradation, extended half-life, and optimized membrane-targeting capabilities [18]. By focusing on these advancements, researchers are developing next-generation AMPs that could revolutionize antibacterial therapy and provide a robust response to the growing threat of antibiotic-resistant infections (Figure 2).
Figure 2: Explaining the agents used against antibiotic targets and resistance where amide linkages can answer both the crisis.
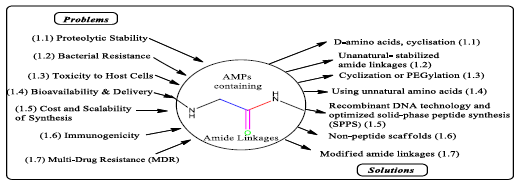
In summary, the need for novel antibacterial agents is urgent, driven by the limitations of existing antibiotics and the rise of resistant pathogens (Figure 3). AMPs represent a promising class of therapeutics with the potential to address these challenges through their unique mechanisms of action and recent advancements in peptide design [19]. Their development is crucial for ensuring effective treatment options in the fight against antibiotic resistance [20].
This review paper highlights the pivotal role of amide linkages in overcoming the limitations of traditional antibiotics and discusses recent progress in AMP design. A thorough understanding of amide linkage dynamics is essential for developing next-generation AMP-based therapeutics, offering a viable solution to the escalating issue of antibiotic resistance.
Figure 3: Illustrations of AMPs bottlenecks and its remedies.
Materials and Methods
Proteolytic stability
One of the primary challenges in the development of Antimicrobial Peptides (AMPs) is their vulnerability to proteolytic degradation. Peptides are often targeted by proteases before they can reach their bacterial targets, reducing their therapeutic effectiveness. Although amide bonds are relatively stable, they can still be susceptible to proteolytic enzymes. To address this, researchers are enhancing peptide stability through modifications such as using D-amino acids, incorporating cyclization, or employing other synthetic strategies to protect peptides from proteolytic attack. discuss engineering plant-derived AMPs for increased stability and resistance to proteolytic degradation, highlighting advances in this area.
Bacterial resistance
The ability of bacteria to develop resistance to AMPs poses another significant challenge. Bacteria can alter membrane compositions, produce proteases, or modify efflux systems to resist AMP action. To counteract these mechanisms, researchers are modifying amide linkages or employing alternative backbone chemistries to create AMPs that are less susceptible to bacterial degradation or resistance. Santos-Junior et al., explore new peptide modifications, including stabilized amide linkages, to overcome bacterial resistance Toward peptide-based approaches to combat multidrug-resistant pathogens.
Toxicity to host cells
Many AMPs disrupt cell membranes in a manner similar to bacterial membranes, which can lead to toxicity in mammalian cells and limit their therapeutic use. To address this, researchers are focusing on stabilizing the amide bond and altering peptide structures through cyclization or PEGylation to enhance selectivity for bacterial membranes over host cells review strategies to reduce toxicity while maintaining antimicrobial potency, emphasizing the need for selective peptide designs. Antimicrobial peptides: Linking partition, activity, and high membrane-bound concentrations.
Bioavailability and delivery
AMPs often suffer from poor bioavailability due to their hydrophilicity and susceptibility to degradation by enzymes in the bloodstream or gastrointestinal tract. Enhancing the pharmacokinetics and bioavailability of AMPs is crucial for effective therapeutic use. Researchers are modifying the amide bond through chemical crosslinking or using unnatural amino acids to improve these properties. Koo and Seo, address these challenges and explore strategies to improve AMP delivery, including peptide modifications for better bioavailability
Cost and scalability of synthesis
The chemical synthesis of AMPs, particularly larger peptides with multiple amide linkages, can be costly and difficult to scale. This limits their practical application in large-scale therapeutic settings. Researchers are exploring new synthesis techniques, including recombinant DNA technology and optimized Solid-Phase Peptide Synthesis (SPPS) methods to reduce production costs. Dijksteel et al., discuss economic and production challenges in scaling up AMPs for clinical trials and potential solutions through synthetic innovations.
Immunogenicity
AMPs with non-natural modifications can sometimes trigger immune responses, limiting their use as long-term therapeutic agents. Researchers are working on modifying the peptide backbone or using non-peptide scaffolds to reduce immunogenicity while preserving antimicrobial function. Haney et al., review immunogenicity issues and how peptide modifications are being used to avoid immune system activation.
Multi-Drug Resistance (MDR)
As bacteria continue to evolve resistance to traditional antibiotics, AMPs are being explored as alternatives. However, there is concern that bacteria may eventually develop resistance to AMPs as well. Researchers are designing AMPs with modified amide linkages to enhance stability and prevent bacterial adaptation. These peptides are often more resistant to bacterial proteases and other resistance mechanisms. Mishra and Wang explore the latest advancements in AMP design, focusing on preventing resistance development.
Clinical translation and regulatory challenges
Despite the promise of AMPs, transitioning from laboratory research to clinical use has been slow due to regulatory approvals and clinical trial challenges. Issues related to peptide stability, scalability, and demonstrating safety have been prominent. Improvements in amide linkage design, better drug delivery systems, and modifications to enhance pharmacokinetics are helping to address these challenges. Mahlapuu et al., discuss the challenges AMPs face in clinical development and regulatory approval, particularly related to stability and safety.
The importance of amide linkages in antibacterial drug discovery
Amide linkages play a pivotal role in the structural integrity and pharmacological efficacy of peptides, which are increasingly explored for their potential as antimicrobial agents in light of escalating antibiotic resistance. Understanding the dynamics of these linkages is crucial for the development of more effective antibacterial therapeutics.
Half-life and therapeutic efficacy in pharmacokinetics
Half-life, specifically elimination half-life, is a crucial concept in pharmacokinetics. It refers to the duration required for the concentration of a drug in the plasma or blood to diminish to half of its initial value. This metric is vital because it provides insight into how quickly a drug is cleared from the body, influencing dosing schedules and treatment plans.
In parallel, therapeutic efficacy denotes a drug's ability to produce the desired beneficial effects. Both half-life and efficacy are critical parameters extensively used in both basic research and clinical settings. The pharmaceutical industry heavily relies on these factors to determine optimal dosages and administration intervals, ensuring that therapeutic effects are maximized while minimizing the risks of toxicity and unwanted accumulation of drugs in various tissues.
The interplay between half-life and therapeutic efficacy is complex. For instance, drugs with short half-lives may not achieve their maximum therapeutic potential, even if administered frequently. Conversely, drugs with very long half-lives run the risk of accumulating in the body, which can lead to toxicity and adverse effects.
This issue is particularly pronounced in the realm of therapeutic peptides. Many of these amide bonds have half-lives of only a few minutes due to factors such as rapid degradation in the body, renal clearance, and low permeability across cell membranes. Chronic diseases like cancer, cardiovascular disorders, and multiple sclerosis often necessitate long-term treatment. In such cases, peptides with extremely short half-lives may prove ineffective, as they struggle to provide consistent therapeutic benefits over extended treatment periods.
Additionally, peptides typically exhibit low oral bioavailability. This limitation arises from their susceptibility to acid hydrolysis and digestive enzymes in the stomach, along with poor absorption in the gastrointestinal tract. While alternative delivery methods such as intravenous or subcutaneous injections can help mitigate some of these absorption issues, the short half-lives of most peptides in the bloodstream, compounded by multiple degradation pathways, restrict their therapeutic utility.
To enhance the effectiveness of peptide drugs and promote their use in clinical settings, it is essential to develop strategies that extend them in vivo half-lives to levels comparable to those of small-molecule drugs with amide linkages. Achieving this goal will not only improve the therapeutic outcomes for patients but also facilitate the broader application of peptide-based therapies in the treatment of various chronic diseases.
Beyond structural stability, amide linkages are instrumental in facilitating the proper folding of peptides into functional secondary structures like alpha-helices and beta-sheets. These conformations are critical for engaging bacterial targets, particularly in membrane-disrupting mechanisms characteristic of many AMPs. Misfolding, often caused by instability in amide linkages, can lead to loss of function, underscoring the necessity of stabilizing this bond.
Biological activity and resistance mechanisms
Antimicrobial Peptides (AMPs) act through diverse mechanisms, such as membrane disruption, which relies on stable peptide structures. The integrity of the amide bond ensures that peptides retain their active conformation, optimizing their interaction with bacterial membranes and intracellular targets. Importantly, modifications of amide bonds, including cyclization and the incorporation of D-amino acids, have been employed to enhance stability and resistance to bacterial proteases.
Peptide linkages, particularly Amide bonds (-CO-NH-), are crucial for the biological activity and structural stability of peptides and proteins. These bonds maintain the integrity of peptide-based drugs, affecting their stability under physiological conditions. Peptide drugs can be susceptible to enzymatic degradation, which limits their therapeutic efficacy. To combat this, researchers have explored modifications to the peptide backbone, such as incorporating non-natural amino acids, to enhance stability and improve drug performance. For example, studies have shown that altering peptide linkages can improve resistance to proteolytic enzymes in peptide therapeutics.
Resistance mechanisms in pathogens, especially bacteria, are a significant challenge for peptide-based drugs, including Antimicrobial Peptides (AMPs). One common resistance mechanism involves the production of proteases by pathogens, which cleave peptide bonds and render the drugs inactive. Enzymes like β-lactamases, which degrade β-lactam antibiotics linked through peptide bonds, are a well-known example of this type of resistance. Additionally, bacteria employ efflux pumps to expel AMPs, reducing the peptides' effectiveness before they can interact with their targets. This resistance mechanism has driven research into modifying peptide linkages to enhance binding affinity for bacterial targets, while reducing efflux pump efficiency.
Another resistance mechanism is the alteration of target sites in bacterial membranes or cell walls, reducing the ability of peptides to bind effectively. For instance, modifications in lipid A, a component of Gram-negative bacterial membranes, can prevent the insertion of AMPs, diminishing their antimicrobial action. Recent strategies to overcome this involve designing peptides that can bind to modified bacterial membranes, addressing resistance while preserving the peptide's biological activity.
Several approaches have emerged to enhance peptide linkages and overcome resistance. Peptide mimetics, which include non-natural peptides designed to mimic the function of natural peptides, have been successful in evading enzymatic degradation. These mimetics, which often feature β-peptides or peptoids, show increased stability and efficacy against drug-resistant pathogens. Cyclization and backbone modifications, such as the introduction of disulphide bridges, have also improved peptide stability and rigidity, making them less susceptible to degradation while enhancing their ability to target pathogens.
Additionally, PEGylation and lipidation strategies, where Polyethylene Glycol (PEG) chains or lipid moieties are attached to peptide linkages, have been effective in protecting peptides from degradation and increasing their half-life in the body. These modifications also improve the peptide's ability to evade resistance mechanisms, such as efflux pumps. Dual-action peptides, which combine multiple mechanisms of action-such as membrane disruption and enzyme inhibition-are another promising approach. These peptides rely on unique peptide linkages that allow them to adopt multiple conformations, increasing their effectiveness against resistant pathogens.
Enhancing stability through chemical modifications
Recent trends in peptide therapeutics focus on enhancing the amide bond's stability via PEGylation, cyclization, and the inclusion of non-natural amino acids. These modifications confer greater resistance to proteolytic degradation, prolong the peptide’s half-life, and improve bioavailability. Such stability is particularly crucial for AMPs, which must endure hostile environments within the host and resist bacterial defence mechanisms.
Cytochrome P450 (CYP) is a large family of proteins that are vital for metabolizing both natural substances produced by living organisms and synthetic drugs created in laboratories or the pharmaceutical industry. In humans, this family includes 18 gene families and 44 subfamilies that code for a variety of CYP enzymes. Most of these enzymes are primarily found in the liver, which is the main site for drug metabolism, although some are also present in other organs like the brain, lungs, and intestines.
Each CYP enzyme has a specific job in processing different drugs. For example, the CYP2C subfamily makes up about 20-25% of all CYP proteins in the liver and is responsible for metabolizing around 15-20% of all medications. This includes commonly used drugs like Non-steroidal Anti-Inflammatory Drugs (NSAIDs), diabetes medications, anticonvulsants, and certain blood pressure medications. On the other hand, the CYP2D6 subfamily is involved in breaking down over 160 different drugs, accounting for about 25% of all prescriptions, including antidepressants, beta-blockers, and opioids. Other CYP families, such as CYP7, CYP17, and CYP27, are involved in processing natural compounds like cholesterol, bile acids, steroids, fatty acids, and vitamin D.
While we understand how important CYP enzymes are for drug metabolism, they seem to play little role in breaking down biological agents, such as proteins. This raises questions about whether small therapeutic peptides with amide linkages which are short chains of amino acids used in treatments are also unaffected by CYP enzymes. More research is needed to explore how these enzymes interact with therapeutic peptides and how this could impact their effectiveness and how the body processes them.
Implications for antimicrobial peptide development
The pursuit of stable amide linkages in peptide design is directly linked to enhanced antimicrobial efficacy. Novel AMPs with modified amide bonds exhibit prolonged activity, reduced toxicity, and increased resistance to bacterial defences, positioning them as leading candidates in next-generation antibiotic therapies. This is particularly pertinent as bacteria evolve mechanisms to inactivate traditional antibiotics, necessitating innovative strategies to maintain therapeutic effectiveness.
Role of amide linkages in antimicrobial activity
Amide linkages are central to the antimicrobial activity of peptides, significantly impacting their structural stability, membrane interactions, and resistance to enzymatic degradation. This detailed exploration highlights how the integrity and modification of amide bonds influence peptide efficacy against bacterial pathogens.
Amide linkages and structural stability: The amide bond is fundamental in maintaining the structural integrity of Antimicrobial Peptides (AMPs). The bond stabilizes secondary and tertiary structures, such as alpha-helices and beta-sheets, which are critical for peptide function. These stable configurations enable AMPs to: Efficiently insert into bacterial membranes. Disrupt membrane integrity by forming pores or channels. Interfere with bacterial enzyme functions through precise binding. The stability provided by amide linkages is crucial for peptides to resist bacterial defences like proteases and maintain prolonged antimicrobial activity.
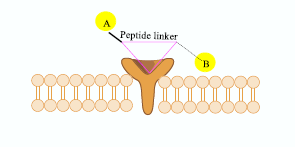
Role of the amide bond in membrane interactions: AMPs often target bacterial membranes, disrupting them through interactions with membrane lipids. The intact amide bonds (Figure 4) help maintain the peptide’s amphipathic nature, which is essential for: Embedding in bacterial cell membranes.
Figure 4: Explains the orientation of the peptide linked spacer.
Creating disruptions in the lipid bilayer leading to cell lysis. Modifications or hydrolysis of the amide bond can alter the peptide’s structure, impacting its amphipathicity and membrane interaction. For instance: Hydrolysed amide bonds can lead to a loss of compact structure, diminishing membrane binding or penetration. Cyclic peptide structures formed via amide linkages often exhibit greater stability and antimicrobial activity due to their rigid configuration. Hydrolysis and modification of amide bonds: Hydrolysis of amide bonds by bacterial enzymes, such as proteases, can deactivate AMPs, reducing their antimicrobial efficacy. Conversely, strategic modifications, such as Nmethylation or cyclization, D-amino acids and unnatural can enhance resistance to enzymatic degradation. These modifications:
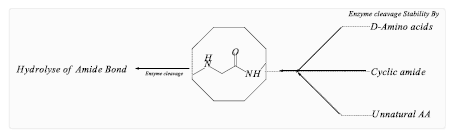
Improve stability against bacterial proteases, extending the peptide’s active lifespan: Increase selectivity for bacterial membranes over host cells, thus reducing toxicity and enhancing therapeutic potential. As shown in the Figure 5. Cyclic peptides, where amide linkages form loops, show improved membrane penetration and are less prone to proteolytic cleavage. Peptides incorporating D-amino acids are less recognizable by bacterial enzymes, enhancing stability and activity against resistant strains.
Figure 5: Showing modifications that are stable by enzyme cleavage.
Enhanced antimicrobial activity through amide modifications: Designing peptides with stable amide linkages can significantly enhance their antimicrobial activity. For instance, cyclic structures with amide linkages often penetrate bacterial membranes more effectively and resist degradation, maintaining a bioactive form longer. Modifications can also tailor peptides to target specific bacterial strains more effectively, particularly in the context of multi-drugresistant bacteria.
Results and Discussion
Mechanisms of action
Antimicrobial Peptides (AMPs) exert their antibacterial effects through various mechanisms primarily centred around their interaction with bacterial cell membranes. The integrity of the amide bond within these peptides plays a crucial role in their effectiveness. For instance, AMPs often disrupt bacterial membranes by inserting into the lipid bilayer, forming pores that lead to cell leakage and death. Peptides with stable amide linkages maintain their structural integrity, which is essential for effective membrane insertion and pore formation. However, bacterial proteases can degrade these peptides, reducing their efficacy. Strategies such as cyclization, incorporating D-amino acids, or using unnatural amino acids can protect amide bonds from proteolysis, thereby enhancing peptide stability.
In the barrel-stave model of membrane disruption, AMPs aggregate to form transmembrane pores. The stability of amide bonds ensures that peptides align correctly for effective pore formation. If the amide bonds are compromised, the peptides may not form pores efficiently. To address this, researchers are introducing non-peptide backbones or cyclizing peptides to preserve structural rigidity and pore-forming capabilities.
The carpet model describes how AMPs coat bacterial membranes, leading to destabilization and membrane fragmentation. Amide bonds contribute to the amphipathic nature of these peptides, aiding their alignment with bacterial membranes. However, bacterial resistance mechanisms, such as altering membrane composition, can reduce peptide efficacy. Modifications like PEGylation or lipidation can enhance targeting and reduce the impact of resistance.
Some AMPs penetrate bacterial membranes to bind with intracellular targets such as DNA or ribosomes, inhibiting vital processes. Maintaining amide bond integrity is crucial for this intracellular targeting. Challenges include degradation by intracellular proteases, which can be mitigated through protease-resistant modifications or small molecule conjugation.
Recent challenges in AMP development include bacterial resistance, peptide stability in biological systems, cytotoxicity to host cells, poor bioavailability, and the difficulty of large-scale synthesis. For example, bacterial resistance can be countered by designing peptides with enhanced stability through amide bond modifications. Stability issues in biological fluids can be improved by incorporating cyclization or D-amino acids. Cytotoxicity can be reduced by modifying amide bonds to increase selectivity for bacterial membranes. To address poor bioavailability, chemical modifications like PEGylation can enhance peptide stability and tissue distribution. Lastly, advances in Solid-Phase Peptide Synthesis (SPPS) and recombinant production have facilitated large-scale synthesis of AMPs.
The amide bond plays a crucial role in the structural integrity and function of Antimicrobial Peptides (AMPs), particularly in maintaining their ability to interact with bacterial membranes and exert antibacterial effects. The stability of the amide bond is essential for preserving the peptide’s structure, ensuring efficient membrane disruption, and facilitating intracellular targeting. Recent challenges in peptide stability, bacterial resistance, cytotoxicity, and large-scale synthesis have led to various solutions involving structural modifications such as cyclization, D-amino acid substitution, and non-peptide backbones, all aimed at enhancing the therapeutic potential of AMPs.
Influence on resistance development
The design and optimization of Antimicrobial Peptides (AMPs) with a focus on amide linkages are crucial for mitigating bacterial resistance. Natural AMPs, which often contain amide bonds, are susceptible to degradation by bacterial proteases that cleave these bonds, rendering the peptides inactive. By incorporating modifications such as D-amino acids or non-natural amino acids, peptides can become more resistant to proteolysis, thereby reducing one potential route for resistance development. This approach ensures that the peptide maintains its activity despite bacterial proteases.
Another strategy involves stabilizing the amide bond to help AMPs resist bacterial membrane modifications. Bacteria may alter their membrane composition to evade peptides that rely on electrostatic interactions. By preserving the peptide's structure through stable amide linkages, peptides can maintain their ability to interact with bacterial membranes even when these membranes are modified. Designing peptides with rigid structures using non-peptide linkers or more stable amide bonds helps ensure activity against resistant bacterial strains.
AMPs may also face resistance through bacterial mechanisms such as efflux pumps or membrane remodelling. Peptides with optimized amide linkages often form more stable structures that are less susceptible to these mechanisms. By integrating stable amide bonds with other modifications, such as cyclization or PEGylation, peptides can better resist bacterial adaptation strategies. This approach makes it more challenging for bacteria to develop resistance.
Furthermore, AMPs targeting bacterial membranes are less likely to induce resistance compared to traditional antibiotics, as altering membrane structure is a complex and costly process for bacteria. Stable amide linkages in AMPs help reduce selective pressure on bacteria, making it less likely for resistance to develop. By maintaining peptide efficacy, stable amide bonds contribute to reducing the likelihood of resistance development.
In conclusion, optimizing amide linkages in AMPs plays a vital role in preventing bacterial resistance. Modifications such as non-natural amino acids, peptide cyclization, and other structural changes help maintain peptide stability and activity. This focus on amide bond chemistry is essential for developing next-generation antimicrobial therapeutics capable of addressing resistant bacterial strains effectively. Advanced computational methods and synthetic strategies are being employed to design novel peptides that leverage the amide bond for enhanced stability and efficacy, highlighting the central importance of these linkages in drug design. developments in peptide design often consider how modifications to the amide linkage can influence the likelihood of bacteria developing resistance. Since many natural antimicrobial peptides utilize amide bonds, optimizing these structures may mitigate the risks associated with resistance.
Conclusion
In conclusion, peptide antibiotics represent a promising frontier in the fight against antibiotic-resistant pathogens, offering broad-spectrum antibacterial activity. However, realizing their full clinical potential requires overcoming significant hurdles, including challenges related to synthesis complexity, proteolytic stability, toxicity, bioavailability, and scalability. The variable efficacy of peptide antibiotics across Gram-positive and Gram-negative bacteria underscores the need for targeted modifications, such as optimizing amide linkages, incorporating non-natural amino acids, and employing peptide cyclization techniques. These innovations not only enhance stability and bioavailability but also minimize the risk of bacterial resistance.
Despite these advancements, the development of peptide-based antibiotics faces ongoing challenges, particularly in ensuring stability against proteolytic degradation and mitigating toxicity and immunogenicity in clinical settings. A deep understanding of Structure-Activity Relationships (SAR) and continuous monitoring of bacterial resistance mechanisms are critical to refining these therapies. Innovations such as peptide mimetics, PEGylation, and the use of alternative backbone chemistries are proving effective in addressing these challenges, providing hope for more resilient peptide therapeutics.
As research progresses, peptide antibiotics are poised to become a vital tool in combating multidrug-resistant infections. Continued efforts to enhance their pharmacokinetics, safety profile, and broad-spectrum activity will be essential to ensuring their successful translation into clinical practice and contributing to the global effort against antibiotic resistance.
References
- Adkins JC, et al. Drugs. 1998;56(6):1055–1064.
[Crossref] [Google Scholar] [PubMed]
- Andersson DI, et al. Nat Rev Microbiol. 2014;12(7):465-478.
[Crossref] [Google Scholar] [PubMed]
- Bengtsson-Palme J, et al. FEMS Microbiol Rev. 2018;42(1):53.
[Crossref] [Google Scholar] [PubMed]
- Blondeau JM. Surv Ophthalmol. 2004;49(2):S73-S78.
[Crossref] [Google Scholar] [PubMed]
- Boissinot J, et al. Front Plant Sci. 2024;15:1381154.
[Crossref] [Google Scholar] [PubMed]
- Brooks BD, et al. Adv Drug Deliv Rev. 2014;78:14-27.
[Crossref] [Google Scholar] [PubMed]
- Bruno BJ, et al. Ther Deliv. 2013;4(11):1443-1467.
[Crossref] [Google Scholar] [PubMed]
- Chambers HF, et al. Nat Rev Microbiol. 2009;7(9):629-641.
[Crossref] [Google Scholar] [PubMed]
- Chiang JY, et al. Liver Res. 2020;4(2):47-63.
[Crossref] [Google Scholar] [PubMed]
- Craik DJ, et al. Chem Biol Drug Des. 2013;81(1):136-147.
[Crossref] [Google Scholar] [PubMed]
- Cruz GS, et al. Antibiotics. 2022;11(12):1772.
[Crossref] [Google Scholar] [PubMed]
- da Cunha NB, et al. Drug Discov Today. 2017;22(2):234-248.
[Crossref] [Google Scholar] [PubMed]
- De La Fuente-Nunez C, et al. Biochim Biophys Acta. 2016;1858(5):1061-1069.
[Crossref] [Google Scholar] [PubMed]
- Dean SN, et al. BMC Microbiol. 2011;11:1-3.
[Crossref] [Google Scholar] [PubMed]
- Domhan C, et al. Molecules. 2019;24(16):2987.
[Crossref] [Google Scholar] [PubMed]
- Drawz SM, et al. Clin Microbiol Rev. 2010;23(1):160-201.
[Crossref] [Google Scholar] [PubMed]
- Feng M, et al. Front Immunol. 2020;11:2030.
[Crossref] [Google Scholar] [PubMed]
- Fosgerau K, et al. Drug Discov Today. 2015;20(1):122-128.
[Crossref] [Google Scholar] [PubMed]
- Furman JL, et al. AAPS J. 2015;17:111-120.
[Crossref] [Google Scholar] [PubMed]
- Gürsoy S, et al. Rheumatol Int. 2008;28:307-311.
[Crossref] [Google Scholar] [PubMed]